EpiGentek
CoviDrop SARS-CoV-2 Specific Furin Cleavage Site (Wild Type) Blocker/Inhibitor Screening Kit
- SKU:
- D-1009
- Availability:
- Usually shipped in 5 working days
- Research Area:
- COVID-19
- Target Applications:
- Amount Quantitation
- Input Type:
- Molecule Compounds
- Vessel Format:
- 96-Well Plate
Description
The CoviDrop™ SARS-CoV-2 Specific Furin Cleavage Site (Wild Type) Blocker/Inhibitor Screening Kit is a complete set of optimized buffers and reagents designed for screening Blockers of SARS-CoV-2 specific furin cleavage site (FCS) and/or inhibitors of furin and other serine proteases such as human air trypsin (HAT) that may also target the FCS of SARS-CoV-2 in a fast and high throughput format.
The kit has the following advantages:
- Fast: Colorimetric assay with easy-to-follow steps for convenience and speed. The entire procedure can be finished within 1 hour and 10 min.
- Robust: Innovative kit composition enables background signals to be extremely low and allows the assay to be simple, accurate, reliable, and consistent.
- Dual-Function: The kit allows for screening of both SARS-CoV-2 FCS direct blockers or furin and furin-like protease activity inhibitors.
- Specific: The substrate contains the entire SARS-CoV-2-S1/S2 cleavage sequence and is proven to be the same as the complete trimeric form of full-length SARS-CoV-2 spike protein in PC cleavage tests. Thus, the assay is uniquely specific for detecting SARS-CoV-2-S1/S2 cleavage and its inhibition by various blockers/inhibitors including small molecule chemicals, therapeutic peptides, antibodies, or other biological inhibitors.
Background
COVID-19 is an infectious disease caused by SARS-CoV-2, a new member of the same coronavirus family that caused SARS and MERS. It was found that the SARS-CoV-2 spike glycoprotein harbors a furin cleavage site (FCS) at the boundary between the S1/S2 subunits, which could be cleaved by furin and/or furin-like PCs secreted from host cells and bacteria in the airway epithelium [1, 2]. Unlike SARS-CoV, cell entry of SARS-CoV-2 is pre-activated by furin and/or furin-like PCs, reducing its dependence on target cell proteases for entry [3]. The cleavage activation of S-protein is well demonstrated to be essential for SARS-CoV-2 spike-mediated viral binding to ACE2, cell-cell fusion, and viral entry into human lung cells [4, 5]. The SARS-CoV-2 furin/PC cleavage site has one core region SPRRAR│SV (eight amino acids, P6–P2). The core region is very unique as its P2 and P3 positions are positively charged residues (Arg), and another residue is hydrophobic (Ala). This status allows such sites to be cleaved by furin or furin-like PC and the cleavage efficiency facilitated by other serine proteases targeting dibasic amino acid sites such as human airway trypsin (HAT).It is well known that SARS-CoV-2 S1/S2 cleavage increases SARS-CoV-2 entry into cells and replication, eventually leading to higher transmission and pathogenicity of COVID-19. Thus, inhibition of SARS-CoV-2 S1/S2 cleavage would allow for the reduction of FCS cleavage-based activation of SARS-CoV-2 spike protein, thereby decreasing viral binding to ACE2, cell-cell fusion, and viral entry into human cells. There are two approaches to block or reduce S1/S2 cleavage of the spike proteins. One approach is based on the inhibition of furin or furin-like protease activity against FCS. For example, peptide or small molecular inhibitors could be used to inhibit furin or furin-like protease activity. Another one is to block SARS-CoV-2 FCS directly. For example, antibody or peptide ligands could be used as blockers against SARS-CoV-2 FCS, avoiding or reducing the disadvantages of inhibiting host furin and furin-like proteases; as such inhibition could non-specifically cause damage to the normal functions of the proteins that need activation by these enzymes. Screening blockers and/or inhibitors of SARS-CoV-2 FCS would help to develop effective drugs for COVID-19 therapy by blocking SARS-CoV-2 cleavage at S1/S2 boundary site.
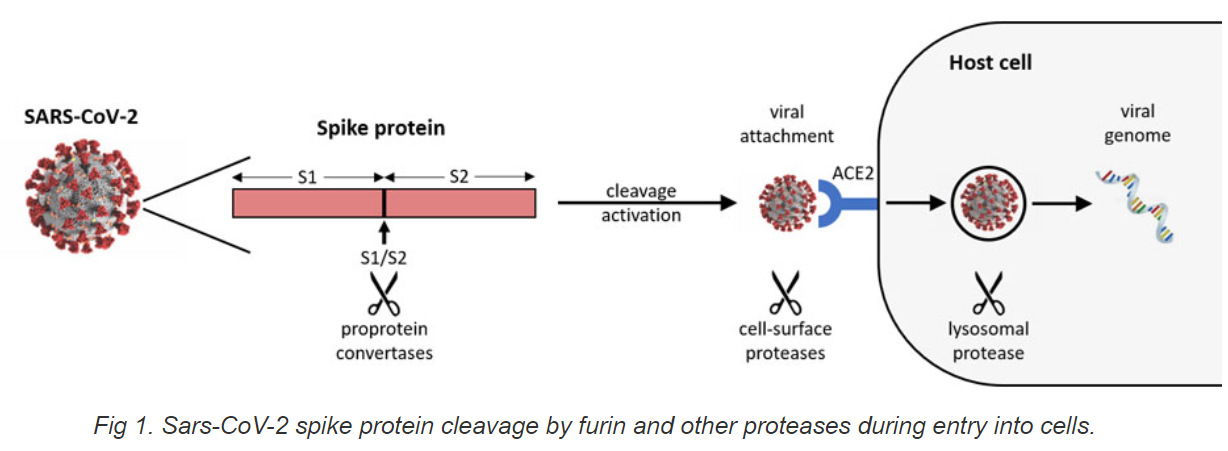
Principles and Procedure
This kit contains all reagents necessary for screening various blockers/inhibitors against SARS-CoV-2-FCS. In this assay, a SARS-CoV-2 specific substrate is tagged with polyhistidine at N-terminal and biotin at C-terminal and bound onto microplate wells through histidine/Ni-NTA at its N-terminal. The cleavage of the substrate at S1/S2 site will remove the S2 part (C-terminal) of the substrate after washing, which causes the decrease in signal generated by avidin/biotin binding after adding Cleavage Detection Solution (CDS). The inhibition of the cleavage by inhibitors will block the reduction of the signals. The signal intensity is measured through an ELISA-like reaction by reading the absorbance in a microplate spectrophotometer at a wavelength of 450 nm. The more the substrate is cleaved, the lower the signal that will be generated. Thus, the PC cleavage activity is inversely proportional to the signal intensity.
Starting Materials
Input materials can be small molecule compounds and other biological molecules that block SARS-CoV-2 specific FSC and/or inhibit furin and serine proteases activity against SARS-CoV-2 specific FCS.